![]()
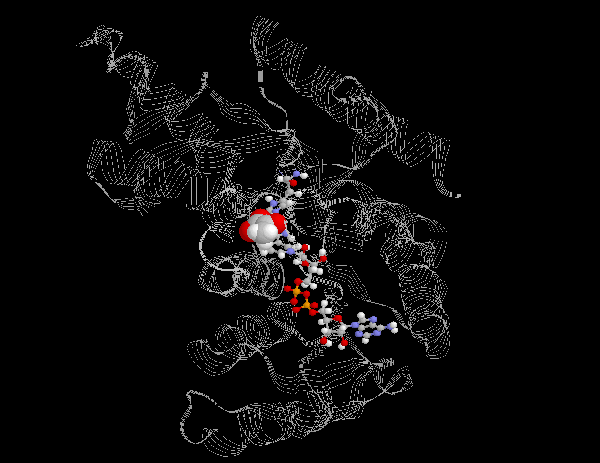
Lactate dehydrogenase catalyzes the reduction of pyruvate to lactate by oxidizing NADH to NAD+. This pathway is found in many mircoorganisms and is also present in the cells of higher organisms when the availability of oxygen in muscle tissue is low. In biochemical terms, lactate is a dead end in metabolism. The reduction of pyruvate to lactate serves only to regenerate NAD+ to allow glycolysis to continue in skeletal muscles. In order for the body to use lactate, it must convert it back into pyruvate through the gluconeogenic pathway in the liver.